Mastering Data Analysis with r read csv for Seamless Data Importation
In the rapidly evolving field of data science, seamless data importation is paramount. This task is often achieved using the ‘read.csv’ function in R. Mastering this function not only simplifies the data analysis workflow but also ensures efficiency in managing large datasets. This article explores practical insights on leveraging’read.csv’ for effective data importation, underpinned by evidence-based statements and real-world examples.
Key Insights
- Primary insight with practical relevance: Utilizing'read.csv' effectively can dramatically reduce data preprocessing time, making it a cornerstone in data analysis.
- Technical consideration with clear application: Understanding and managing various parameters within'read.csv' is crucial for optimal data importation and analysis.
- Actionable recommendation: Routinely practice 'read.csv' variations with different data formats to ensure mastery.
Efficiency through Standard Usage
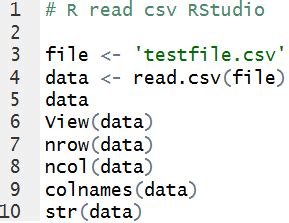
‘Read.csv’ is an indispensable function for importing comma-separated values files into R. Its straightforward syntax provides default settings that often work well for many datasets. However, the true power of’read.csv’ lies in its adaptability. By customizing its parameters, one can significantly enhance the efficiency of the data importation process. For example, settingstringsAsFactors = FALSE prevents automatic conversion of character data to factors, thus preserving the original data type and avoiding unintended data transformations.
Advanced Customization Techniques
Beyond the standard usage,‘read.csv’ offers an array of parameters for tailored data importation. For instance, usingheader = TRUE indicates that the first row contains variable names, while na.strings = "" allows for specifying how missing values are represented in the dataset. These parameters allow for detailed control over the data importation process, crucial when dealing with inconsistently formatted datasets. Another important parameter, fill = TRUE, ensures that the function fills missing values with blanks, helping in maintaining data completeness during the import.
Consider a dataset where different files use different conventions for missing values; setting `na.strings = c("NA", "", "?")` can handle these variations effectively. For large datasets, using `nrows` can limit the number of rows read in, which is essential for avoiding memory overflow during the import process. An example implementation might look like this:
data <- read.csv("large_dataset.csv", na.strings = c("NA", "", "?"), nrows = 1000)
This approach ensures that only a manageable subset of the data is read into memory, facilitating smoother data analysis.
What if my CSV file has different delimiters?
In such cases, use the `sep` parameter to specify the correct delimiter. For instance, for a tab-separated file, set `sep = "\t". If your file uses a semicolon, use `sep = ";"`. This parameter tells R how to parse your data correctly.
How can 'read.csv' handle UTF-8 encoded files?
To handle UTF-8 encoded files, use the `fileEncoding` parameter. For example, when reading an encoded file, you would use `read.csv("data.csv", fileEncoding = "UTF-8")`. This ensures that characters are correctly interpreted during the import process.
By mastering ‘read.csv’, data analysts can streamline their workflow, saving valuable time and resources. The ability to efficiently import data is foundational to any successful data analysis project. Regular practice and exploration of ‘read.csv’ variations will enhance your proficiency, making it an indispensable tool in your data science toolkit.